SIMSPINE: A Biomechanics-Aware Simulation Framework for 3D Spine Motion Annotation and Benchmarking
Biomechanics-aware simulation, vertebra-level 3D annotations, and reference baselines for spine motion estimation from RGB.
The SIMSPINE dataset and source code are publicly available. Inference support is provided through the SpinePose Inference Library; SimSpine inference models will be added there soon.
Frames
2.14M
Spine Keypoints
15
Subjects / Actions
7 / 15
Markers / Axes
37 / 62
SIMSPINE represents a step toward scalable, biomechanics-aware spine motion estimation from RGB video.
Abstract
Modeling spinal motion is fundamental to understanding human biomechanics, yet remains underexplored in computer vision due to the spine’s complex multi-joint kinematics and the lack of large-scale 3D annotations. We present a biomechanics-aware keypoint simulation framework that augments existing human pose datasets with anatomically consistent 3D spinal keypoints derived from musculoskeletal modeling. Using this framework, we create the first open dataset, named SIMSPINE, which provides sparse vertebra-level 3D spinal annotations for natural full-body motions in indoor multi-camera capture without external restraints. With 2.14 million frames, this enables data-driven learning of vertebral kinematics from subtle posture variations and bridges the gap between musculoskeletal simulation and computer vision. In addition, we release pretrained baselines covering fine-tuned 2D detectors, monocular 3D pose lifting models, and multi-view reconstruction pipelines, establishing a unified benchmark for biomechanically valid spine motion estimation. Specifically, our 2D spine baselines improve the state-of-the-art from 0.63 to 0.80 AUC in controlled environments, and from 0.91 to 0.93 AP for in-the-wild spine tracking. Together, the simulation framework and SIMSPINE dataset advance research in vision-based biomechanics, motion analysis, and digital human modeling by enabling reproducible, anatomically grounded 3D spine estimation under natural conditions.
Highlights
First open RGB benchmark in this setting to pair vertebra-level 3D spine positions with rotational kinematics for natural full-body motion.
Simulation pipeline combines multi-view spinal detection, Human3.6M body markers, OpenSim inverse kinematics, and virtual vertebral markers.
Reference baselines span 2D spine detection, multi-view 3D triangulation, and monocular 3D lifting, with strong gains from consistent simulation-derived supervision.
Overview
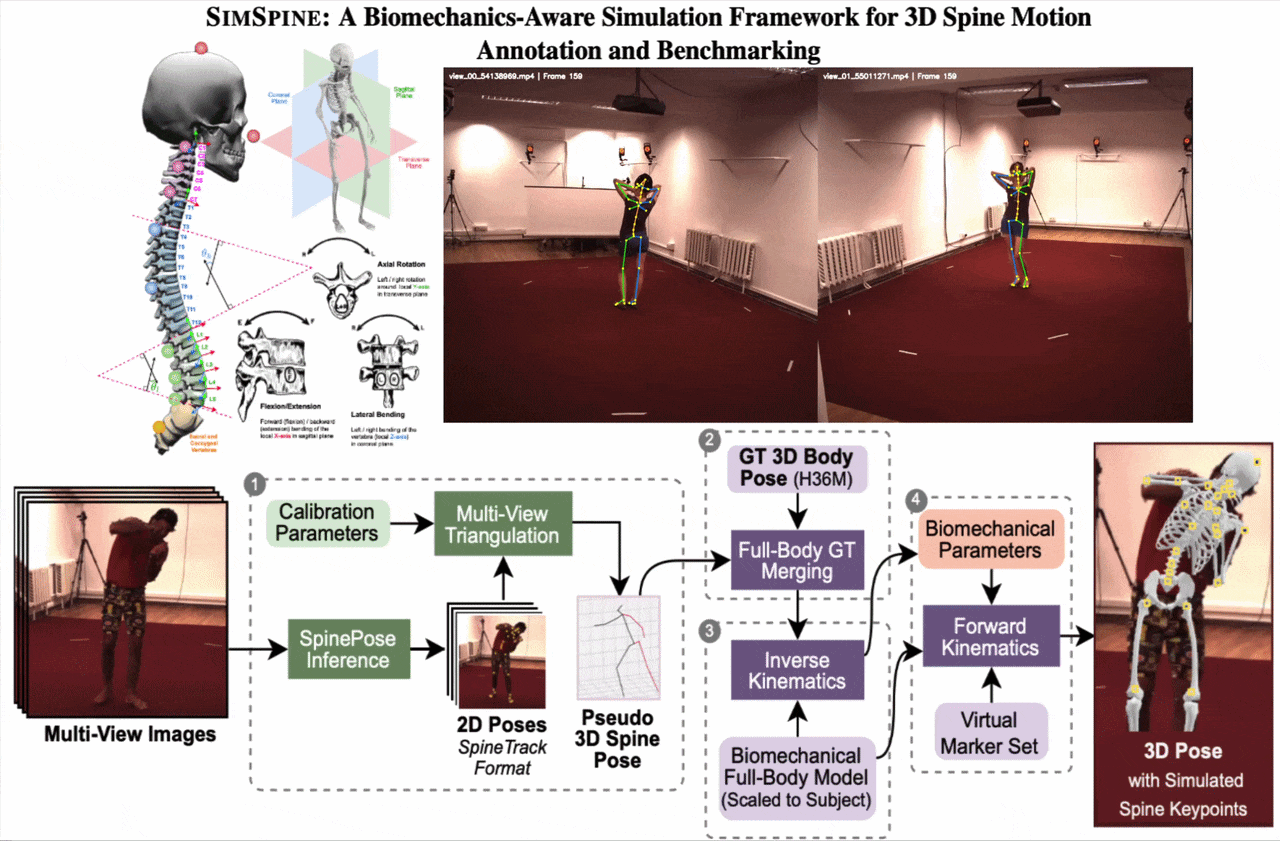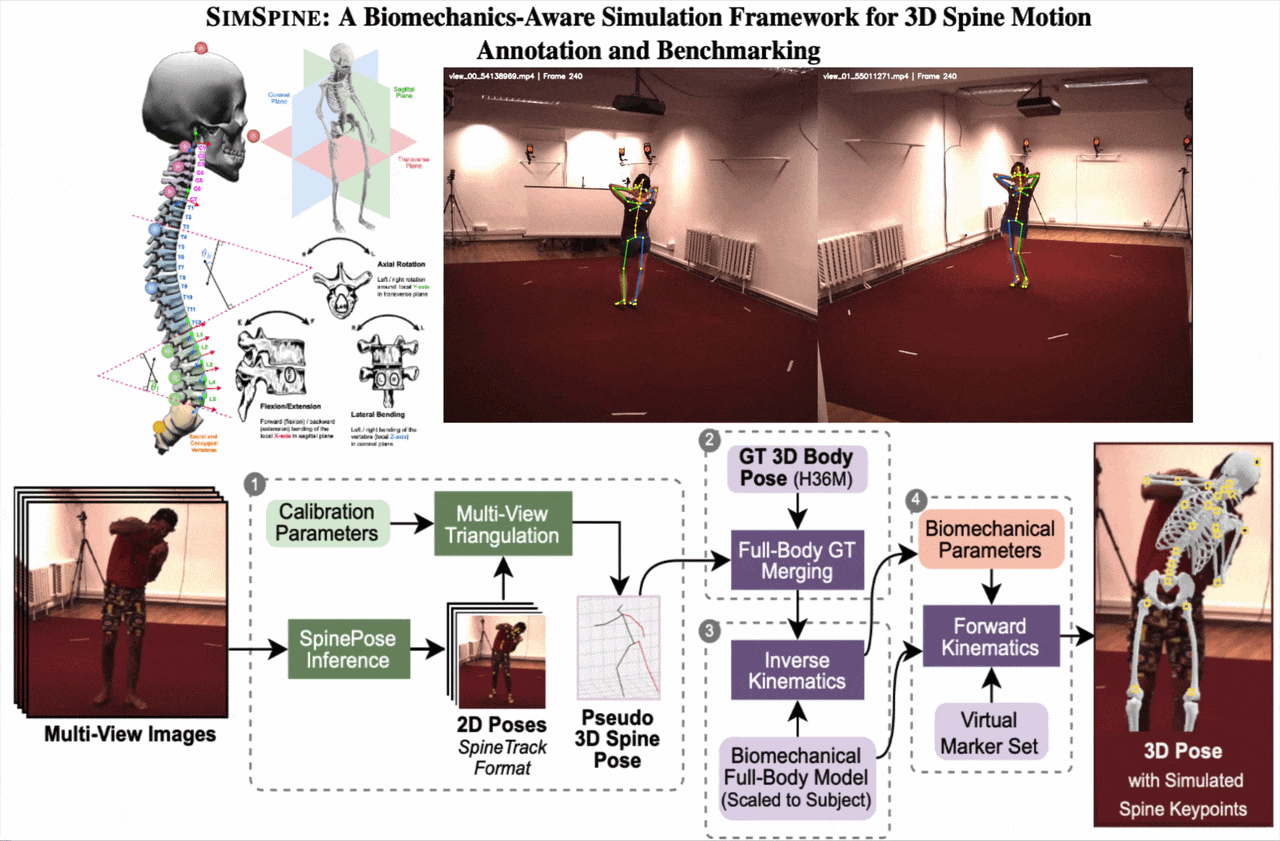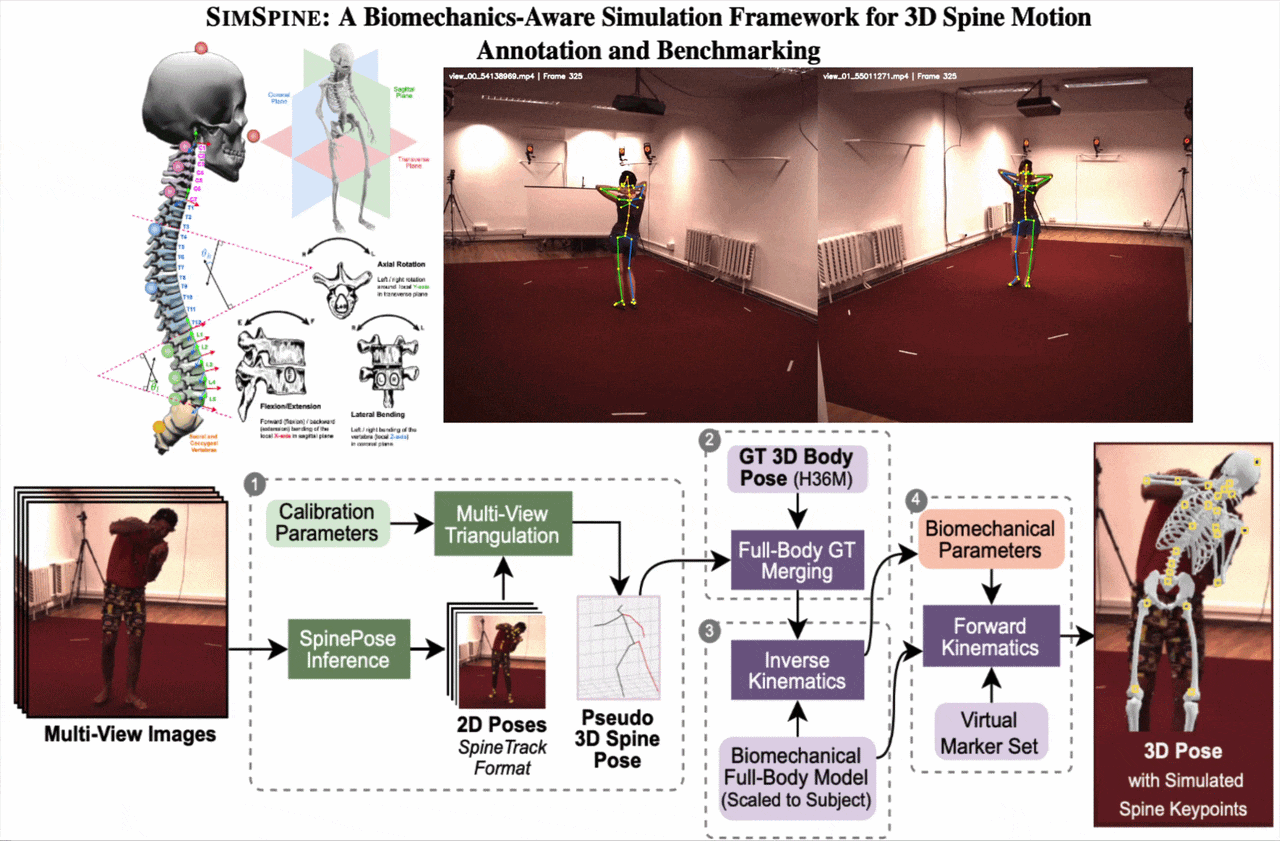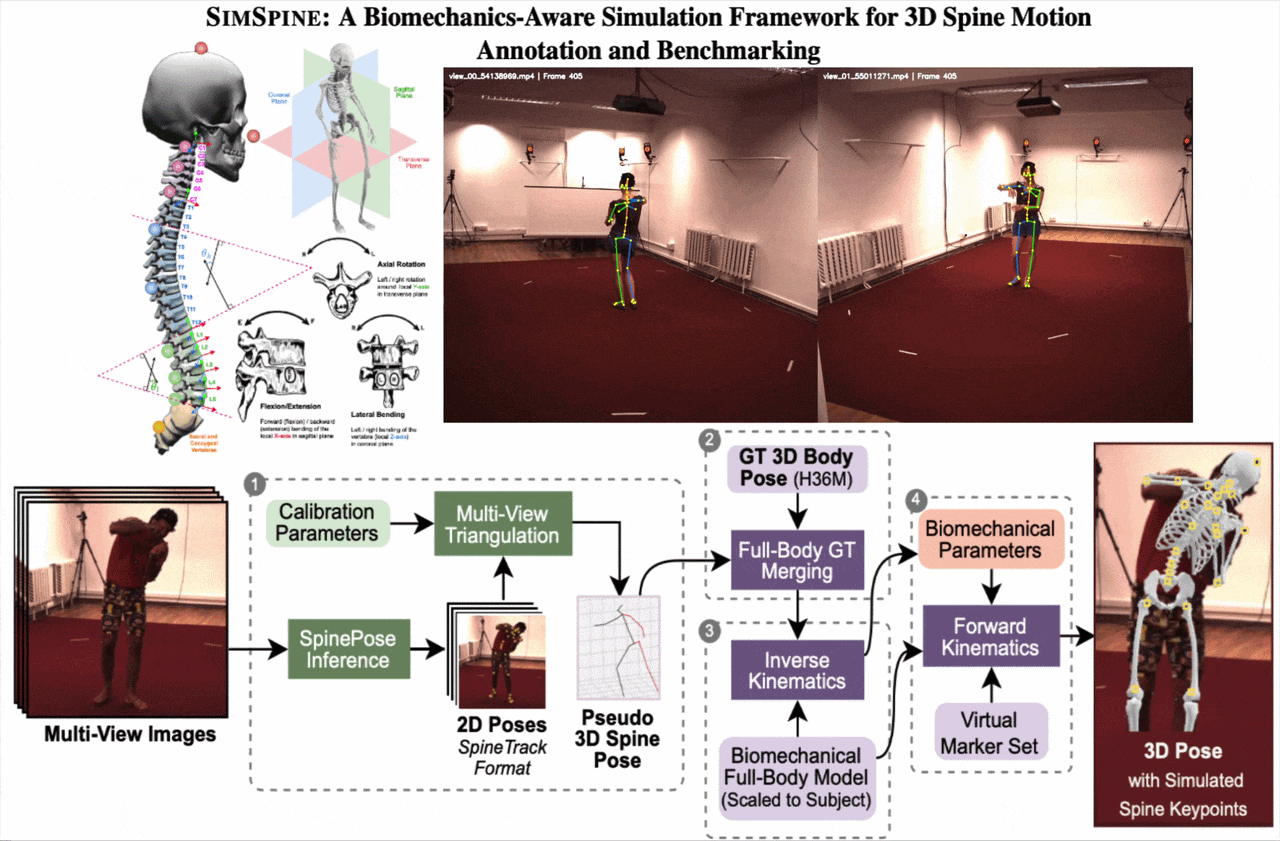
SIMSPINE turns a conventional multi-view whole-body motion dataset into a spine-aware benchmark. The key idea is to infer sparse spinal observations from RGB, combine them with the native Human3.6M body markers, fit a subject-scaled musculoskeletal model, and then export anatomically constrained vertebral keypoints and rotations through forward kinematics.
The project addresses a persistent gap in computer vision: most pose datasets capture limbs well, but they do not expose vertebra-level motion that matters for biomechanics, ergonomics, rehabilitation, and digital human modeling. SIMSPINE closes part of that gap by providing a scalable annotation pipeline and a benchmark dataset that remain grounded in anatomical structure rather than unconstrained pose-only priors.
The resulting benchmark covers three core tasks. First, 2D spine keypoint estimation from RGB. Second, multi-view 3D reconstruction in metric world coordinates. Third, monocular 2D-to-3D lifting for root-relative spine pose. Together these baselines make the page useful not only as a paper summary, but also as a reference for future experiments.
What this project page covers
- A compact view of the annotation pipeline and its modeling assumptions.
- A dataset snapshot with the main counts, splits, and annotation types.
- Biomechanical sanity checks for curvature and lumbar/cervical motion.
- Reference results for 2D, multi-view 3D, and monocular 3D baselines.
- Direct links to the dataset, paper, and code resources.
Method
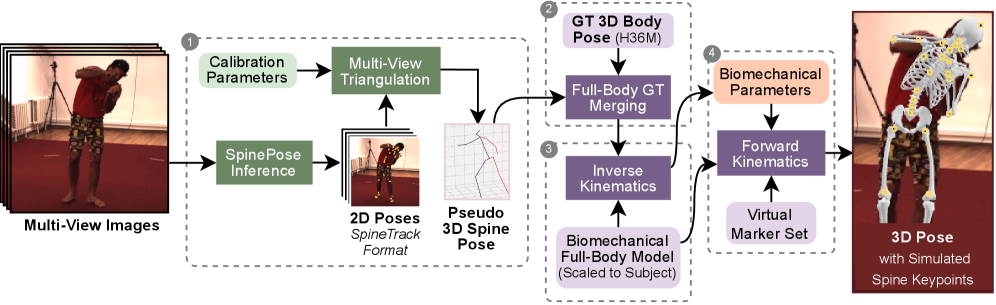
From synchronized RGB views, SIMSPINE triangulates spine detections, merges them with Human3.6M body markers, fits a subject-scaled OpenSim model with inverse kinematics, and recovers vertebral keypoints and biomechanical parameters through forward kinematics.
1. Multi-view spinal pseudo-labels
A pretrained 2D spine detector predicts spinal landmarks in each synchronized view. Robust triangulation, view-consistency checks, reprojection filtering, and low-pass temporal smoothing produce pseudo-3D spinal observations.
2. Merge with Human3.6M markers
The pseudo-3D spine points are aligned with the native Human3.6M 3D markers. This gives a single marker set that contains both reliable body supervision and spine-specific observations.
3. Subject-scaled inverse kinematics
An OpenSim model derived from Rajagopal et al. with lumbar refinements is scaled to each subject. Inverse kinematics solves a weighted least-squares fitting problem over time with smoothness regularization.
4. Virtual vertebral markers
Virtual markers attached to vertebral bodies are exported through forward kinematics. The pipeline also outputs Euler-angle rotations around anatomical axes for downstream regression and analysis.
5. Quality control and modeling assumptions
Frames with implausible curvature are removed, rare angular discontinuities are clamped, and short gaps are interpolated to maintain continuity. The lumbar region is fully articulated from T12-L1 through L5-S1 with three rotational degrees of freedom per segment, while the cervicothoracic region is represented by one aggregate 3-DOF joint. Thoracic and remaining cervical bodies are kept rigid to preserve identifiability from RGB inputs.
Dataset and annotation space
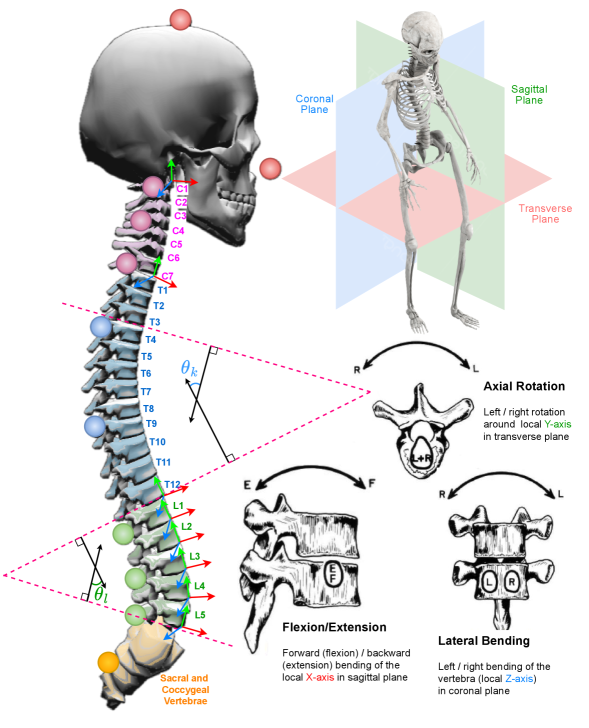
The spine model is divided into cervical, thoracic, and lumbar regions and exposes spine-centric landmarks together with vertebral rotations around anatomical axes.
Dataset snapshot
| Total frames | 2.14M |
|---|---|
| Train / test split | 1.56M train, 0.58M test |
| Subjects / actions | 7 subjects, 15 actions |
| Train subjects | S1, S5, S6, S7, S8 |
| Test subjects | S9, S11 |
| Spine-centric landmarks | 15 total: 9 along the vertebral column, 2 skull landmarks, 2 clavicle landmarks, and 2 shoulder-blade landmarks |
| Marker set | 37 markers total: 12 Human3.6M limb points, 15 new spine points, and 10 pseudo-labels on feet and face |
| Kinematic outputs | 62 axes, including 56 Euler angles |
| Labels | 3D vertebral positions and per-segment rotational kinematics |
Compared with prior public resources, the paper positions SIMSPINE as the first open benchmark in this space that connects RGB video to vertebra-level 3D kinematics. The benchmark is built on Human3.6M, so the visual domain is indoor multi-camera motion capture with licensed source data rather than in-the-wild pathology imagery.
Biomechanical validity checks
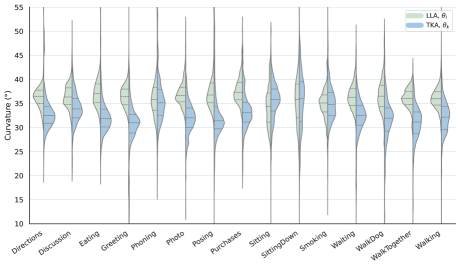
Action-conditioned thoracolumbar curvature remains within reported adult ranges. Across actions, lumbar lordosis is centered around roughly 32-41° and thoracic kyphosis around 28-39°.
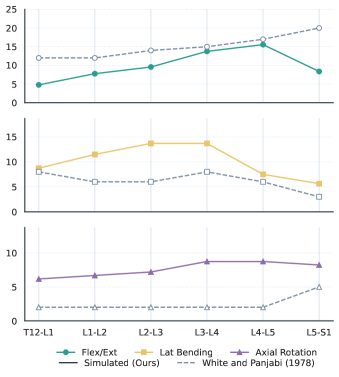
The lumbar motion trends follow expected qualitative patterns: flexion/extension increases toward the caudal lumbar levels, lateral bending peaks in the mid-lumbar region, and axial rotation stays relatively small.
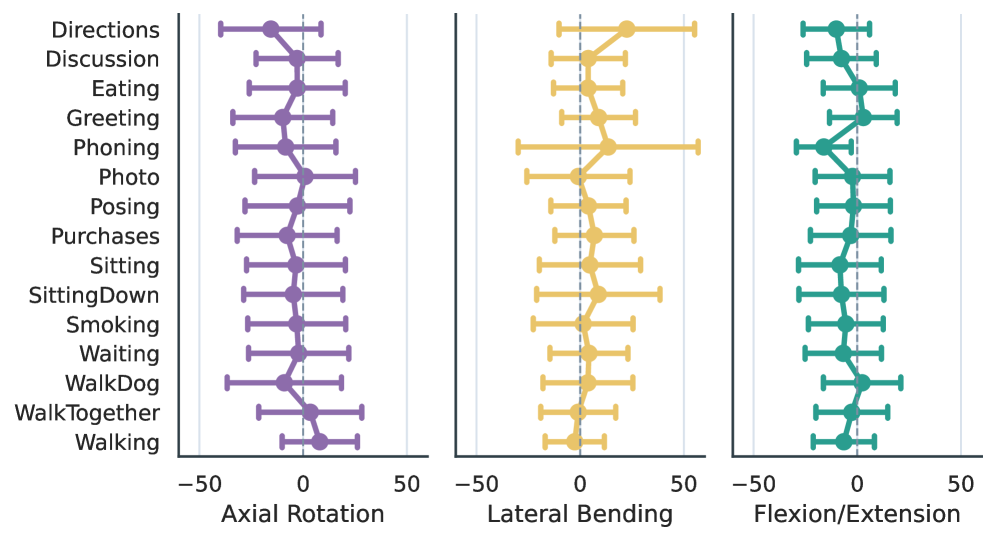
The aggregate cervical proxy stays close to neutral while still reflecting task-dependent head motion. Actions such as talking on the phone show larger lateral-bending spread than neutral standing or walking actions.
Interpretation
These checks matter because the paper does not claim direct in-vivo measurement. Instead, it shows that simulation-derived annotations remain anatomically plausible, action-sensitive, and consistent with known lumbar trends. That makes the benchmark suitable as supervision for learning and benchmarking, while still keeping the right boundary around clinical use.
Reference benchmarks
Top-line results
| Task | Setting | Result |
|---|---|---|
| 2D pose estimation | Best indoor result on SimSpine | 0.803 AUC with SpinePose-l-ft |
| 2D pose estimation | Best outdoor spine tracking on SpineTrack | 0.928 APS / 0.937 ARS with SpinePose-m-ft |
| Multi-view 3D reconstruction | Fine-tuned 2D detections | 31.82 mm MPJPE and 29.53 mm P-MPJPE on the full skeleton |
| Multi-view 3D reconstruction | Oracle setting with GT 2D | 7.85 mm MPJPE and 1.79 mm P-MPJPE on the full skeleton |
| Monocular 3D lifting | Detected 2D, full-body training | 16.28 mm P-MPJPE on spine joints |
| Monocular 3D lifting | GT 2D, full-body training | 13.48 mm P-MPJPE and 25.94 mm MPJPE on spine joints |
2D detection
Consistent simulation-derived supervision materially improves the controlled indoor benchmark and still transfers to outdoor spine tracking. The strongest paper-level summary is the jump from 0.63 to 0.80 AUC indoors and from 0.91 to 0.93 APS outdoors.
Multi-view 3D
Triangulation quality is highly tied to 2D detection quality. Fine-tuning the 2D detector reduces the overall MPJPE from 49.30 mm to 31.82 mm, showing that better spine labels are immediately useful for downstream geometry.
Monocular 3D lifting
Training on the full-body joint set improves over a spine-only input representation. The gain is visible both for detected 2D input and in the GT 2D upper-bound setting, which suggests that global context helps vertebral localization.
Ablation takeaway. Only a small fraction of indoor simulated data is needed. The paper reports that using 2% of SIMSPINE, about 31k indoor images, already gives near-saturated performance, and per-batch mixing is the most stable way to balance indoor and outdoor supervision.
Resources
Paper and project links
PDF, arXiv abstract, Hugging Face dataset, inference library, and source repository.
Dataset
SIMSPINE is publicly available on Hugging Face: dfki-av/simspine.
Model weights
Inference-ready SpinePose models are currently available via the SpinePose Inference Library. SimSpine model support will be added there in an upcoming release.
Evaluation code
Benchmark instructions, configs, and evaluation scripts for 2D detection, 3D lifting, and multi-view reconstruction will be maintained in the public codebase.
Demo video / talk
A talk recording and demo media will be linked here after the presentation materials are finalized.
Licensing note
The paper states that code, models, and SIMSPINE spine annotations will be released for research use. Full-body keypoints remain reproducible through the pipeline on licensed Human3.6M data.
Limitations and intended use
- The benchmark is simulation-derived and should be treated as a scalable proxy for method development, not as direct clinical measurement.
- The lumbar spine is detailed, but the thoracic and cervical regions are simplified for identifiability from RGB.
- Intervertebral translations, rib-cage coupling, and force-consistent dynamics are not modeled.
- The visual domain comes from Human3.6M, so appearance diversity and pathology coverage are limited.
- The most appropriate use is pretraining, benchmarking, and controlled development before fine-tuning on smaller biomechanically validated datasets.
Acknowledgement
This work was co-funded by the European Union’s Horizon Europe research and innovation programme under Grant Agreement No. 101135724 (LUMINOUS) and Grant Agreement No. 101092889 (SHARESPACE).
BibTeX
@inproceedings{khan2026simspine,
title={SIMSPINE: A Biomechanics-Aware Simulation Framework for 3D Spine Motion Annotation and Benchmarking},
author={Khan, Muhammad Saif Ullah and Stricker, Didier},
booktitle={Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR)},
month={June},
year={2026},
pages={}
}Maintained by saifkhichi96 on GitHub.
The website is distributed under different open-source licenses. For more details, see the notice at the bottom of the page.